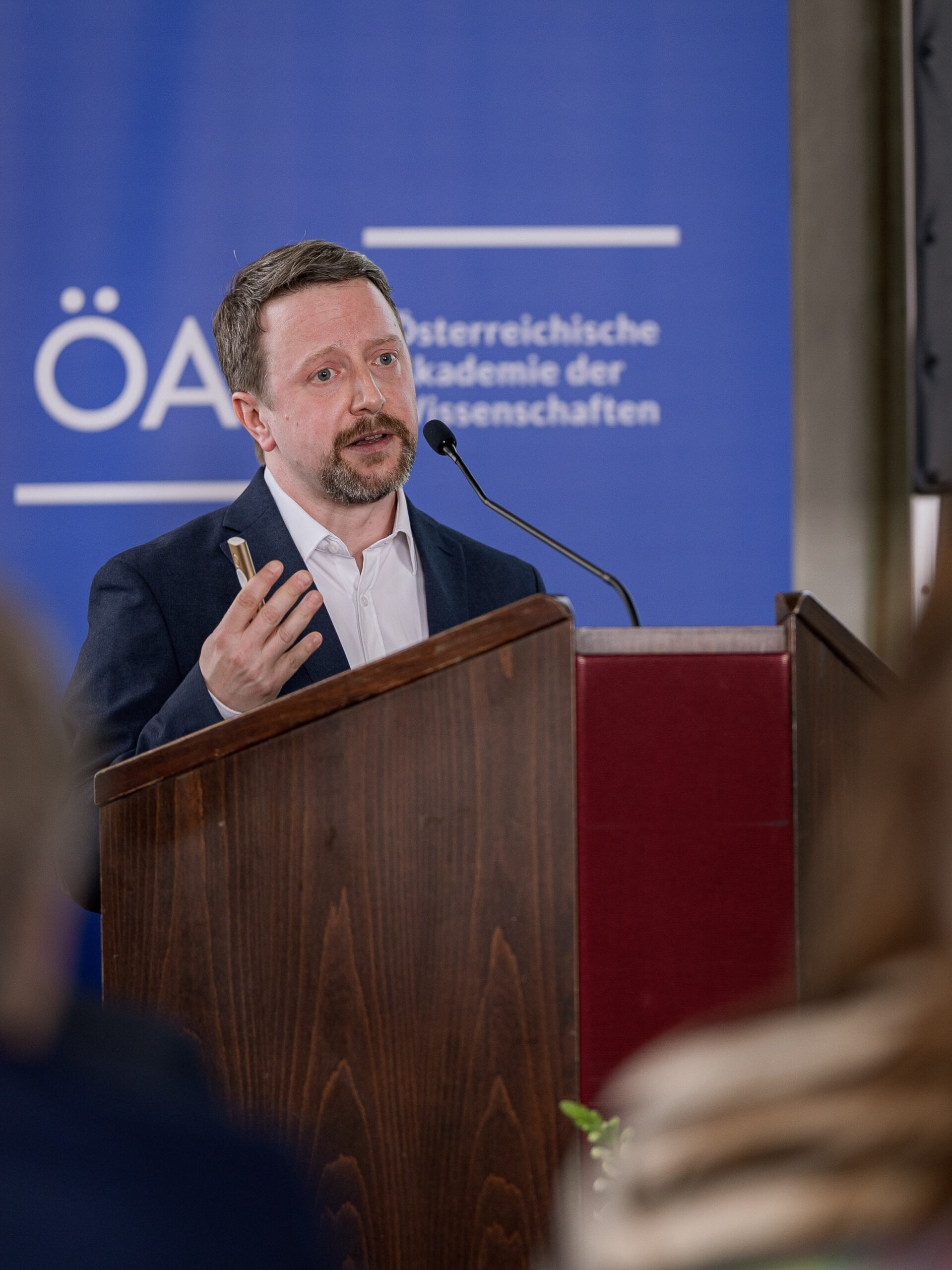
Florian Halbritter awarded Johann Wilhelm Ritter von Mannagetta Prize for Medicine

A computational scientist turned cancer researcher, Florian Halbritter is uncovering how developmental errors drive childhood cancer—work that has now earned him the Johann Wilhelm Ritter von Mannagetta Prize for Medicine from the Austrian Academy of Sciences (ÖAW). The award recognizes scientists under 45 working in Austria who have made outstanding contributions to cancer research.
Halbritter, principal investigator at St. Anna CCRI, is recognized for his outstanding contributions to understanding how pediatric cancers arise—and how they might be treated more precisely in the future.
From machine learning to cancer research
Halbritter’s path into biology was anything but conventional. He studied cognitive science at the University of Osnabrück in Germany, focusing on computer science and machine learning. However, he soon became interested in applying his skillset to answer biological questions, and he joined the Institute for Stem Cell Research at the University of Edinburgh, where he obtained a PhD in Computational Biology.
“I initially knew very little about biology,” Halbritter admits. “But I found it fascinating. I learned a lot from my colleagues and supervisor, and I used the computational tools I knew to study gene regulatory networks in stem cells.”
Halbritter’s work, which focused on how new cell identities are established, led him to join Christoph Bock’s lab at CeMM as a postdoc. There, he first came into contact with pediatric cancer research through a collaboration with Caroline Hutter’s group at St. Anna CCRI. “Working with them, I learned that many childhood cancers arise from errors in cell differentiation,” Halbritter recalls. “It was also my first time interacting with clinician scientists, and I was inspired by their drive to make discoveries with real impact for patients.”
The experience was transformative for Halbritter, who decided to join St. Anna CCRI to pursue a new mission. “Working with the Hutter lab, we had used computational biology to analyze Langerhans Cell Histiocytosis tumors,” he explains. “We realized they were made up of many different cell types—but we could only study them once they had already formed. I wanted to understand how they develop in the first place.”
Understanding where development goes wrong – and how to fix it
In 2018, Halbritter established his own research group at the institute, merging his unique background in developmental biology, genomics and computational analysis to study how mistakes in normal development cause pediatric cancer.
By comparing healthy and cancer-prone development in stem cell models, his team studies how mutations rewire gene regulatory networks and prevent cells from maturing properly—a key feature of pediatric cancer.
The team’s ultimate goal is to identify new molecular targets for innovative therapeutic approaches. “Targeting the disease-causing mutation is an obvious choice, but doesn’t always work,” Halbritter explains. “By understanding how whole molecular pathways within the cell are affected, we might be able to find other ways to prevent or undo the effects of the mutations.”
This work contributes to a central goal in pediatric oncology: developing treatments that are more targeted and effective. “Many current treatments work great for some high-risk cancer types but fail in others,” Halbritter says. “In those cases, finding new, specific therapeutic targets is essential to give patients hope for a cure.”
About the Johann Wilhelm Ritter von Mannagetta Prize for Medicine
The Johann Wilhelm Ritter von Mannagetta Prize for Medicine, awarded annually by the Austrian Academy of Sciences (ÖAW), recognizes scientists under 45 working in Austria who have made outstanding contributions to cancer research.
“I’m very thankful for this recognition,” Halbritter says. “Coming from a different background, it is very special.”
He emphasizes that the award reflects a collective effort: “Along the way, I’ve had the fortune of meeting many great scientists who shaped the direction of my career and kindly introduced me to their research fields,” he reminisces. “All members of my research team over the years, as well as our collaborators within St. Anna CCRI and beyond, have been essential to this success.”